Content
The projects that you can currently find here are:
- Climate change and temperature anomalies
- General Social Survey (GSS)

In the following, you can have a quick look at some projects that I am currently working on. As these are not yet finished, you may find missing explanations, interpretations or planned but not yet executed analyses.
I know you’re excited to see more of my work but please bear with me! These are works in progress and once I was able to continue, refine, finish and polish these, they will appear as individual projects in my portfolio for you to inspect.
Consider this as a teaser… they will leave you wanting more!
The projects that you can currently find here are:

We want to study climate change and find data for that on the Combined Land-Surface Air and Sea-Surface Water Temperature Anomalies in the Northern Hemisphere at NASA’s Goddard Institute for Space Studies. The tabular data of temperature anomalies can be found here
To define temperature anomalies you need to have a reference, or base, period which NASA clearly states that it is the period between 1951-1980.
First, we load the file:
weather <- read_csv("https://data.giss.nasa.gov/gistemp/tabledata_v3/NH.Ts+dSST.csv",
skip = 1,
na = "***")Here, we use two additional options: skip and na.
skip=1 option is there as the real data table only starts in Row 2, so we need to skip one row.na = "***" option informs R how missing observations in the spreadsheet are coded. When looking at the spreadsheet, you can see that missing data is coded as "***". It is best to specify this here, as otherwise some of the data is not recognized as numeric data.Once the data is loaded, we notice that there is a object titled weather in the Environment panel. We inspect the dataframe by clicking on the weather object and looking at the dataframe that pops up on a seperate tab.
For each month and year, the dataframe shows the deviation of temperature from the normal (expected). Further, the dataframe is in wide format.
Before we dive into the data, we want to transform it to a more helpful format.
The weather dataframe has a column for Year and then one column per month of the year (12 more in total). However, there are six further columns that we will not need. In the code below, we use the select() function to select the 13 columns (year and the 12 months) of interest to get rid of the others (J-D, D-N, DJF, etc.).
We also convert the dataframe from wide to ‘long’ format by using the pivot_longer() function. We name the new dataframe tidyweather, the variable containing the name of the month month, and the temperature deviation values delta.
tidyweather <- weather %>%
select(Year:Dec) %>%
pivot_longer(cols = 2:13, names_to = "Month", values_to = "delta")
glimpse(tidyweather)## Rows: 1,680
## Columns: 3
## $ Year <dbl> 1880, 1880, 1880, 1880, 1880, 1880, 1880, 1880, 1880, 1880, 188…
## $ Month <chr> "Jan", "Feb", "Mar", "Apr", "May", "Jun", "Jul", "Aug", "Sep", …
## $ delta <dbl> -0.54, -0.38, -0.26, -0.37, -0.11, -0.22, -0.23, -0.24, -0.26, …When inspecting the dataframe with the glimpse() function or by opening the separate tidyweather dataframe tab, we find that our dataset has been reduced to the following three variables now:
Year,Month, anddelta, or temperature deviation.Let us plot the data using a time-series scatter plot, and add a trendline. To do that, we first create a new variable called date in order to ensure that the delta values are plotted chronologically.
We now have a Month variable that includes the months “Jan”, “Feb”, etc. as characters and a month variable that includes those months as ordered factors, i.e. “Jan”<“Feb”< etc.
#create new variable `date` to ensure chronological order
tidyweather <- tidyweather %>%
mutate(date = ymd(paste(as.character(Year), Month, "1")),
month = month(date, label=TRUE),
year = year(date))
#plot time-series scatter plot with time variable on x-axis and temp deviation on y-axis
ggplot(tidyweather, aes(x=date, y = delta))+
geom_point()+
geom_smooth(color="red") + #add red trend line
labs (title = "Increasing weather anomalies in the past few decades",
subtitle = "Temperature deviations per month over time",
x = "Year",
y = "Temp deviation from expectation"
)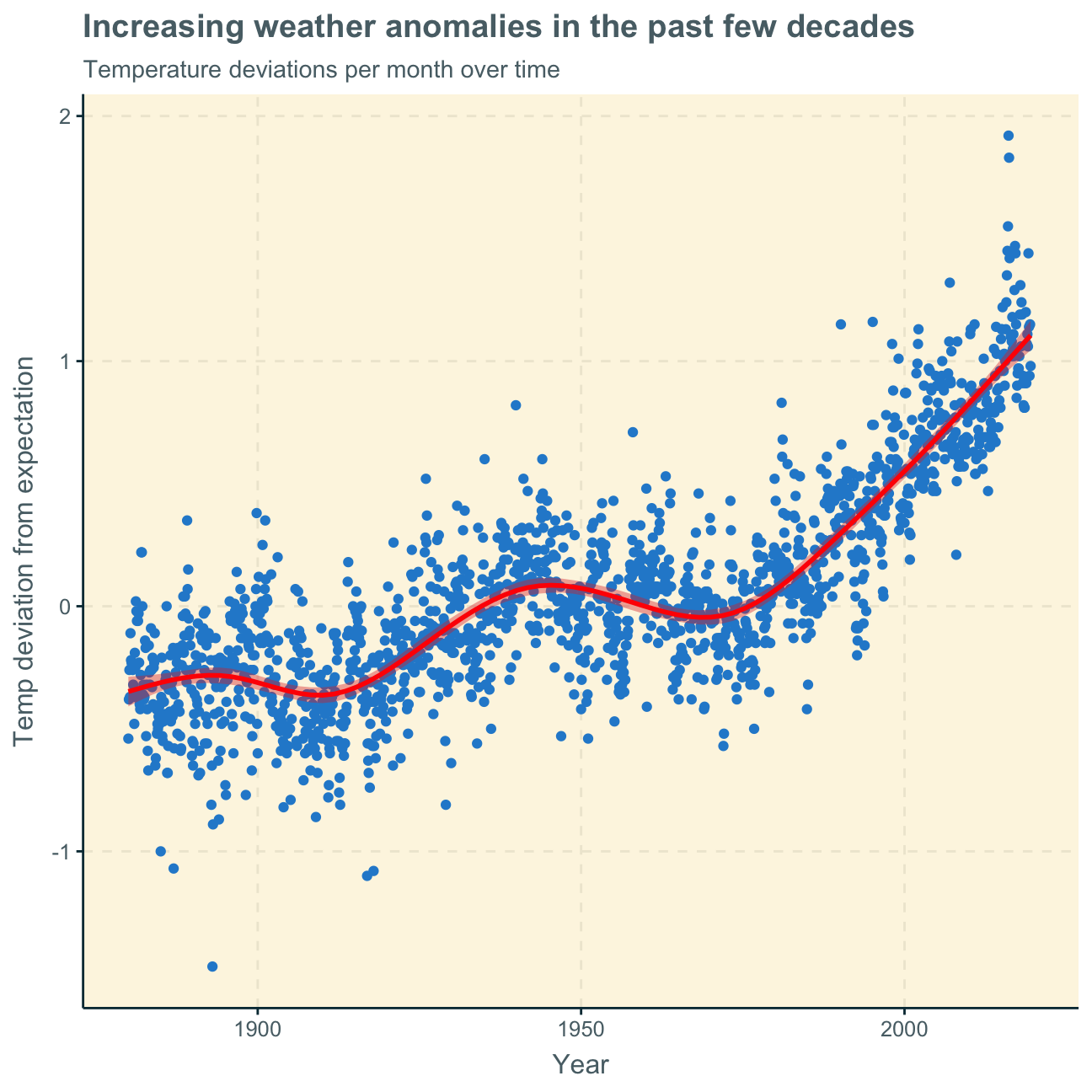
Next, we want to find out if the effect of increasing temperature deviations is more pronounced in some months. We use facet_wrap() to produce a separate scatter plot for each month, again with a smoothing line.
ggplot(tidyweather, aes(x=date, y = delta))+
geom_point()+
geom_smooth(color="red") +
#facet by month
facet_wrap(~month) +
labs (title = "Temperature's rising!",
subtitle = "Temperature deviations per month over the years",
x = "Year",
y = "Temp deviation from expectation"
)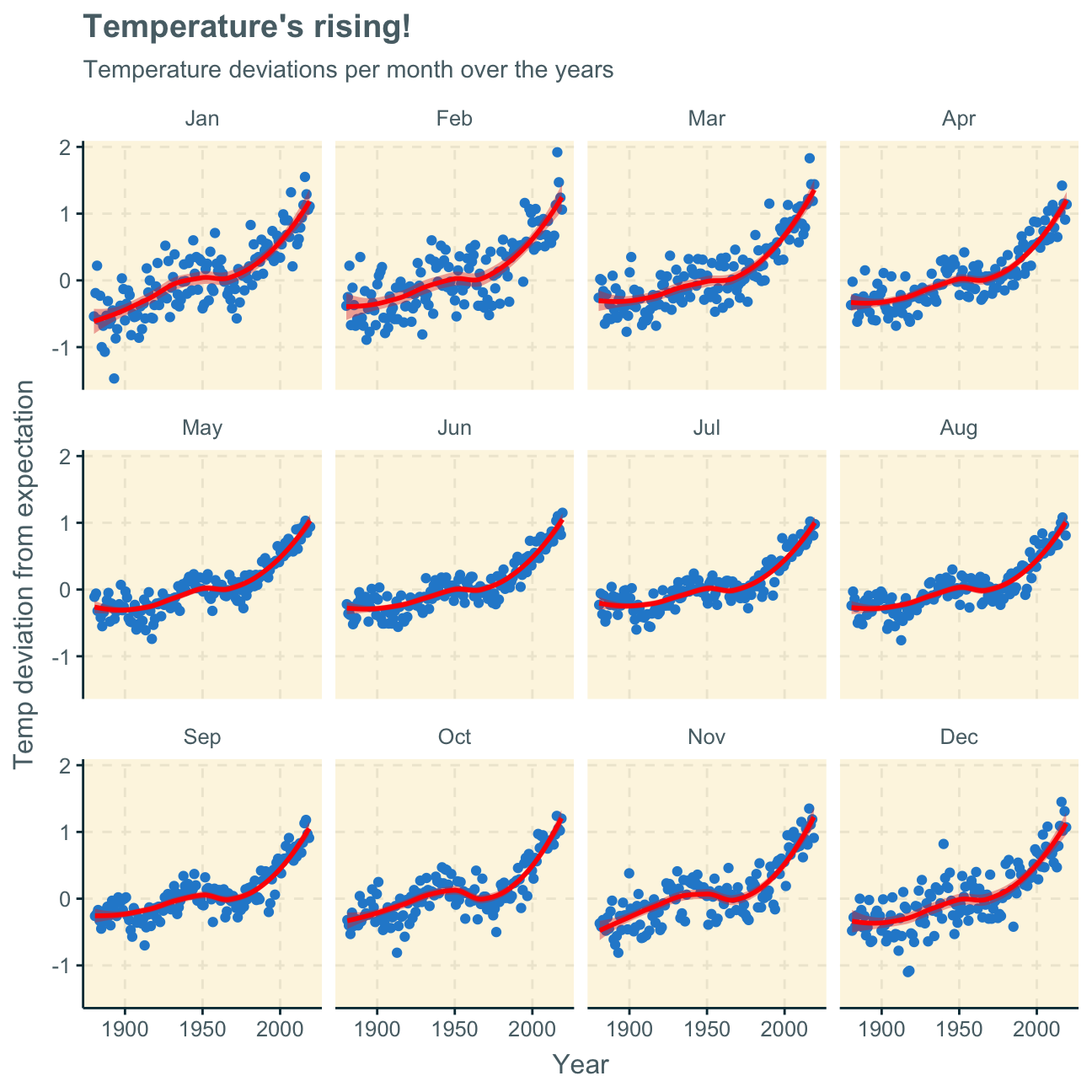
Looking at the produced plots, we find that the increase in temperature deviations from the normal (expected) temperature has increased over the years in every month of the year. Although the trend line has similar shapes in all months, the curve is flatter in some months and steeper in others. We find that the delta has increased more significantly in the winter months (e.g. Nov, Dec, Jan) than the summer months (e.g. Jun, Jul, Aug). This may mean that winters have become significantly hotter or colder but summers have only become slightly hotter or colder over the past decades.
To investigate the historical data further, it may be useful to group it into different time periods.
NASA calculates a temperature anomaly, as difference form the base period of 1951-1980. The code below creates a new data frame called comparison that groups data in five time periods: 1881-1920, 1921-1950, 1951-1980, 1981-2010 and 2011-present.
We remove data before 1800 and before using filter. Then, we use the mutate function to create a new variable interval which contains information on which period each observation belongs to. We assign the different periods using case_when().
comparison <- tidyweather %>%
filter(Year>= 1881) %>% #remove years prior to 1881
#create new variable 'interval', and assign values based on criteria below:
mutate(interval = case_when(
Year %in% c(1881:1920) ~ "1881-1920",
Year %in% c(1921:1950) ~ "1921-1950",
Year %in% c(1951:1980) ~ "1951-1980",
Year %in% c(1981:2010) ~ "1981-2010",
TRUE ~ "2011-present"
))Inspecting the comparison dataframe in the Environment pane, we find that the new column interval has been added and shows which period each observation belongs to.
Now that we have the interval variable, we can create a density plot to study the distribution of monthly deviations (delta), grouped by the different time periods we are interested in. We set fill to interval in order to group and colour the data by different time periods.
ggplot(comparison, aes(x=delta, fill=interval))+
geom_density(alpha=0.2) + #density plot with transparency set to 20%
#theme
labs (
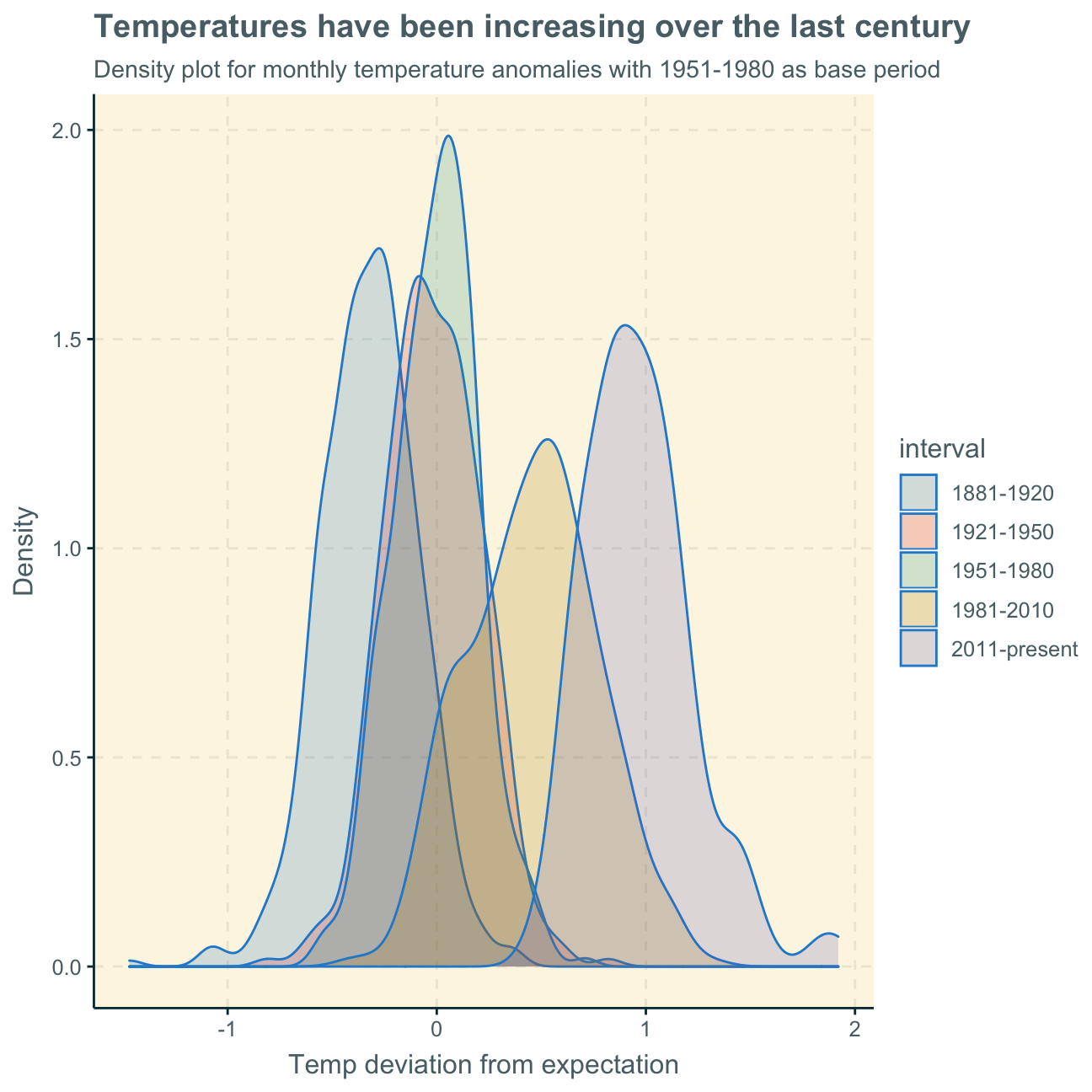
title = "Temperatures have been increasing over the last century",
subtitle = "Density plot for monthly temperature anomalies with 1951-1980 as base period",
y = "Density", #changing y-axis label to sentence case
x = "Temp deviation from expectation"
)
So far, we have been working with monthly anomalies. However, we are also interested in average annual anomalies. We do this by using group_by() and summarise(), followed by a scatter plot to display the result.
#creating yearly average delta
average_annual_anomaly <- tidyweather %>%
group_by(Year) %>% #grouping data by Year
# creating summaries for mean delta
# use `na.rm=TRUE` to eliminate NA (not available) values
summarise(annual_average_delta = mean(delta, na.rm=TRUE))
#plotting the data:
ggplot(average_annual_anomaly, aes(x=Year, y= annual_average_delta))+
geom_point()+
#Fit the best fit line, using LOESS method
geom_smooth(method = "loess") +
#change to theme_bw() to have white background + black frame around plot
labs (
title = "Significant temperature anomalies since 1970s",
subtitle = "Average yearly temperature deviation from the normal",
y = "Average annual temp anomaly",
x = "Year"
) 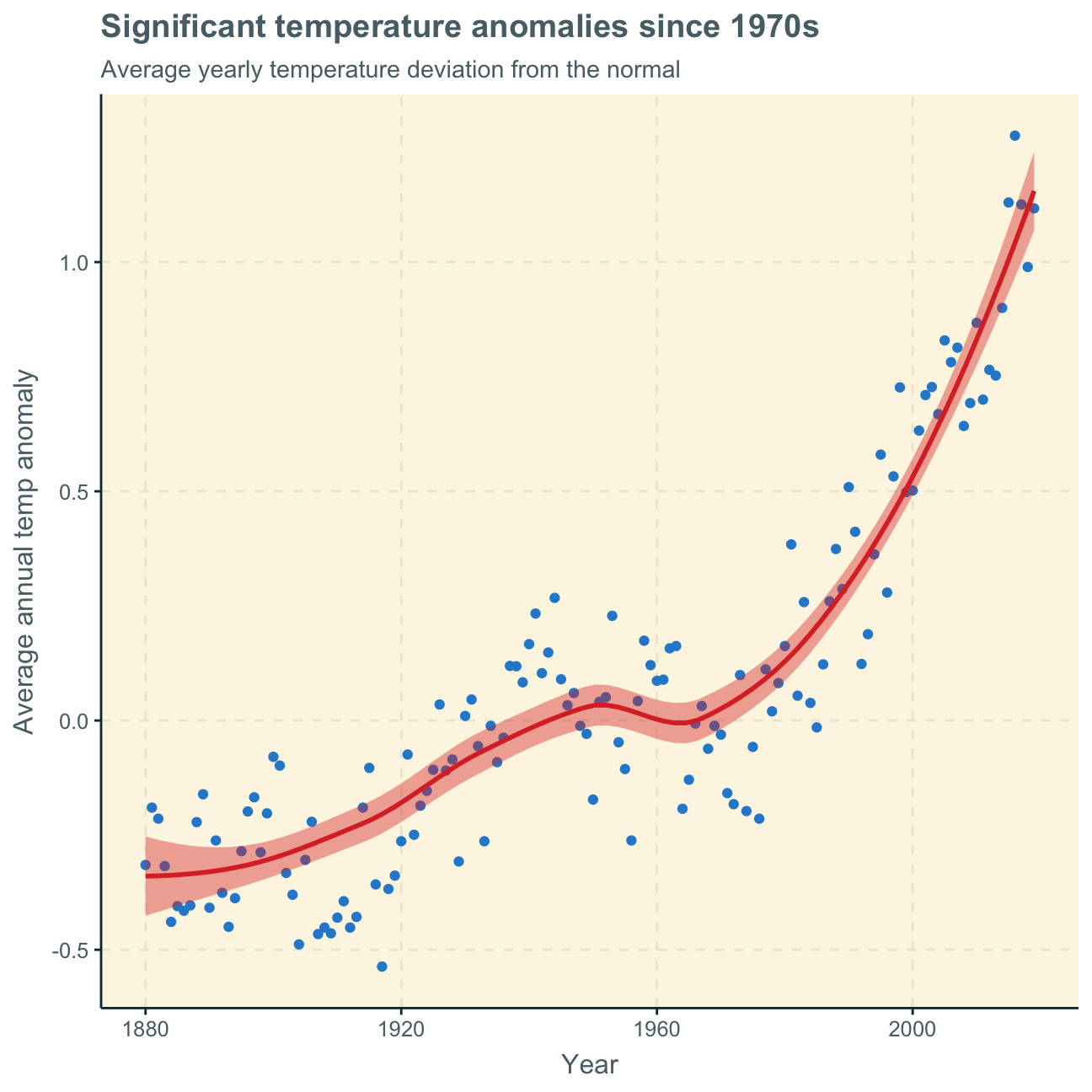
The analyses of monthly and annual temperature anomalies show very similar results. Over time, i.e. over both months and years, temperature overall increases. As the base period for the “normal” is 1951-1980, it is obvious that the deviations from the expected temperature in these years are relatively small (close to zero). This results in the small stagnating part of the curve around these years. The deviations before that time period are negative with the greatest negative deviation at the beginning of the observation years in 1880, decreasing the negative with every year after. After the base period, the deviations become positive and increasingly higher. The positive deviations are especially steep after around 1980.
This graph clearly shows an average temperature increase over the past century since 1880 with especially significant increases in the past few decades since 1980. This clearly depicts what is commonly known as climate change and how rising temperatures have been fueled by technologies and lifestyle of the 20th and 21st century.
In a next step, it would be interesting to split the data into geographical sections instead of periodical sections to investigate which specific regions of the world are more or less affected by climate change.
deltaNASA points out on their website that
A one-degree global change is significant because it takes a vast amount of heat to warm all the oceans, atmosphere, and land by that much. In the past, a one- to two-degree drop was all it took to plunge the Earth into the Little Ice Age.
Here, we will construct a confidence interval (CI) for the average annual delta since 2011. We use the dataframe comparison as it has already grouped temperature anomalies according to time intervals and we are only interested in what is happening between 2011-present.
First, we construct the CI by using a formula.
formula_ci <- comparison %>%
filter(interval == "2011-present") %>% #choose the interval 2011-present
# calculate summary statistics for temperature deviation (delta)
# calculate mean, SD, count, SE, lower/upper 95% CI
summarise(mean = mean(delta, na.rm = TRUE), SD = sd(delta, na.rm = TRUE), count = n(), SE = SD/sqrt(count), ci_lower = mean - 1.96*SE, ci_upper = mean + 1.96*SE)
#print out formula_CI
formula_ci## # A tibble: 1 x 6
## mean SD count SE ci_lower ci_upper
## <dbl> <dbl> <int> <dbl> <dbl> <dbl>
## 1 0.966 0.262 108 0.0252 0.916 1.02#CI = [0.916, 1.02]Second, we construct the CI by using a bootstrap simulation with the infer package.
library(infer)
#set seed number
set.seed(1234)
boot_temp <- comparison %>%
# Select 2011-present
filter(interval == "2011-present") %>%
# Specify the variable of interest
specify(response = delta) %>%
# Generate a bunch of bootstrap samples
generate(reps = 1000, type = "bootstrap") %>%
# Find the mean of each sample
calculate(stat = "mean")
#calculate 95% confidence interval
percentile_ci <- boot_temp %>%
get_ci(level = 0.95, type = "percentile")
percentile_ci #print 95% CI## # A tibble: 1 x 2
## lower_ci upper_ci
## <dbl> <dbl>
## 1 0.917 1.02Using formulas and individually calculating the mean, standard deviation, count and standard error gives us the results of the last but one code chunk above. We construct a 95% CI by both adding and subtracting the standard error times 1.96 (z score for 95% confidence) to/from the mean. We find the CI to be [0.916, 1.02].
Using the bootstrap simulation in the last code chunk, we generate multiple bootstrap samples and find the mean of each of these samples to then find that the 95% CI is [0.917, 1.02].
When using the summary statistics, we are 95% confident that the average yearly temperature anomaly is between [0.916, 1.02]. When using bootstrap, we found this interval to be [0.917, 1.02].
These results support our earlier analysis. The confidence intervals show us that a positive temperature deviation of around 1 degree Celcius is highly likely to occur, proving the overall hypothesis that our temperatures are increasing every year (by around 1 degree Celcius). As define by the NASA earlier, these seemingly small temperature increases can have significant implications for the Earth, our climate and our nature.